exercises:2016_ethz_mmm:md_slab
Hot gold
TO USE THE FUNCTION LIBRARY (VERSION UP TO DATE) IN THE INTERACTIVE SHELL:
you@eulerX ~$ module load courses mmm vmd
you@eulerX ~$ mmm-init
This exercise deals with heating a gold slab, namely the (100) reconstructed that you already simulated last time. The goal is to plot a density profile in the direction orthogonal to the slab, and to compute (using vmd) the radial distribution function g( r ) at various temperatures.
Download the 4.2 exercise into your $HOME folder and unzip it:
you@eulerX ~$ wget http://www.cp2k.org/_media/exercises:2016_ethz_mmm:exercise_4.2.zip you@eulerX ~$ unzip exercises:2016_ethz_mmm:exercise_4.2.zip you@eulerX ~$ cd exercise_4.2
All files of this exercise ( input and scripts are all commented) can be also downloaded from the wiki: exercise_4.2.zip
- First, we simulate the system at 700 K using a thermostat.
you@eulerX exercise_4.2$ bsub cp2k.popt -i 700.inp -o 700.out
- Then, the obtained xyz trajectory can be analyzed using the script histo_z available in the directory.
you@eulerX exercise_4.2$ ./histo_z 700-pos-1.xyz
The output is 700-pos-1.xyz.z, a file with three columns: z, dn/dz, and the progressive integral of this quantity.
Assignments:
- Explain the profile, and use the third column to draw conclusions about the surface structure.
- Study the source of the script. Understand its behavior.
- Copy histo_z into another file and modify it to only include the particles from the first 10 frames of the trajectory.
- Run it and see the differences to the first profile.
- Do the same excluding the first 10 frames.
- Explain those differences, based on what you see in the *.ener file (energies, temperature…).
- Perform consequently a simulation at T=1100 K and T=1300 K (files: 1100.inp and 1300.inp):
you@eulerX exercise_4.2$ bsub cp2k.popt -i 1100.inp -o 1100.out you@eulerX exercise_4.2$ bsub cp2k.popt -i 1300.inp -o 1300.out
- And again analyze these trajectories using the script histo_z:
you@eulerX exercise_4.2$ ./histo_z 1100-1-pos.xyz you@eulerX exercise_4.2$ ./histo_z 1300-1-pos.xyz
Assignments:
- Discuss the differences in the density profile. What do you expect to see in vmd?
- Now, use vmd to look at the trajectories. As you launch vmd,
in Tk console you can:
Load a pbc.vmd file which includes the definition of the periodic box
vmd> source pbc.vmd
Draw the box:
vmd> draw pbcbox
Wrap all atoms in the periodic box:
vmd> pbc wrap -first first -last last
- Try to play with representations: color the surface atoms in one color, the bulk ones in another color.
- Using the “radial distribution function” plugin from the extension menu (Extensions>Analysis>Radial Pair Distribution Function g( r ) ), draw the g( r ) of the system.
Assignments:
- Discuss radial distribution function for 700, 1100, and 1300 K.
Hint: how to use the g( r ) module:
- First apply pbcs (see above)
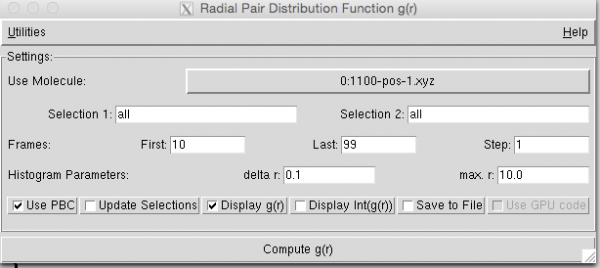
- Open the radial distribution function plugin and enter the parameters as shown (note: in the example below we excluded the first 10 frames) (from “Utilities” you can check that your unit cell is OK)
- Click “Compute g( r )”
- From the “File” menu of the graph window, you can save as postscript file or other formats.
exercises/2016_ethz_mmm/md_slab.txt · Last modified: by 127.0.0.1