Table of Contents
Basic electronic structure calculation
In this exercise, you will perform a first basic electronic structure calculation to obtain the molecular orbitals (MOs) of Ethene: Your calculations will produce a list of occupied and non occupied MOs and a series of *.cube files, that allow the visualization of the orbitals with VMD.
1. Step: Run the calculation
Create a new directory for this exercise and run a CP2K calculation with the following (commented) input file:
- ethene.inp
&GLOBAL PROJECT ethene RUN_TYPE ENERGY PRINT_LEVEL MEDIUM &END GLOBAL &FORCE_EVAL METHOD Quickstep ! Electronic structure method (DFT,...) &DFT &PRINT &MO_CUBES ! Controls which MOs are written to cube-files. NHOMO 5 NLUMO 5 &END MO_CUBES &END PRINT &POISSON ! Solver requested for non periodic calculations PERIODIC NONE PSOLVER WAVELET ! Type of solver &END POISSON &QS ! Parameters needed to set up the Quickstep framework METHOD GAPW ! Method: gaussian and augmented plane waves &END QS &SCF ! Parameters controlling the convergence of the scf. This section should not be changed. MAX_ITER_LUMOS 10000 EPS_SCF 1.0E-6 SCF_GUESS ATOMIC MAX_SCF 60 EPS_LUMOS 0.000001 &OUTER_SCF EPS_SCF 1.0E-6 MAX_SCF 6 &END &END SCF &XC ! Parametes needed to compute the electronic exchange potential &XC_FUNCTIONAL NONE ! No xc functional &END XC_FUNCTIONAL &HF ! Hartree Fock exchange. In this case is 100% (no fraction specified). &SCREENING ! Screening of the electronic repulsion up to the given threshold. EPS_SCHWARZ 1.0E-10 ! Threshold specification &END SCREENING &END HF &END XC &END DFT &SUBSYS &CELL ABC 10 10 10 PERIODIC NONE ! Non periodic calculations. That's why the POISSON section is needed &END CELL &TOPOLOGY ! Section used to center the atomic coordinates in the given box. Useful for big molecules &CENTER_COORDINATES &END &END &COORD C -2.15324 3.98235 0.00126 C -0.83403 4.16252 -0.00140 H -0.25355 3.95641 0.89185 H -0.33362 4.51626 -0.89682 H -2.65364 3.62861 0.89669 H -2.73371 4.18846 -0.89198 &END COORD &KIND H ! Basis set and potential for H &BASIS 2 1 0 0 3 1 18.73113700 0.03349460 2.82539370 0.23472695 0.64012170 0.81375733 1 0 0 1 1 0.16127780 1.00000000 &END POTENTIAL ALL &POTENTIAL 1 0 0 0.20000000 0 &END &END KIND &KIND C ! Basis set and potential for C &BASIS 4 1 0 0 6 1 3047.52490000 0.00183470 457.36951000 0.01403730 103.94869000 0.06884260 29.21015500 0.23218440 9.28666300 0.46794130 3.16392700 0.36231200 1 0 1 3 1 1 7.86827240 -0.11933240 0.06899910 1.88128850 -0.16085420 0.31642400 0.54424930 1.14345640 0.74430830 1 0 1 1 1 1 0.16871440 1.00000000 1.00000000 1 2 2 1 1 0.80000000 1.00000000 &END POTENTIAL ALL &POTENTIAL 4 2 0 0.34883045 0 &END &END KIND &END SUBSYS &END FORCE_EVAL
2. Step
If the calculation was performed correctly, a number of new files should have been written:
$ ls *.cube ethene-WFN_00004_1-1_0.cube ethene-WFN_00008_1-1_0.cube ethene-WFN_00012_1-1_0.cube ethene-WFN_00005_1-1_0.cube ethene-WFN_00009_1-1_0.cube ethene-WFN_00013_1-1_0.cube ethene-WFN_00006_1-1_0.cube ethene-WFN_00010_1-1_0.cube ethene-WFN_00007_1-1_0.cube ethene-WFN_00011_1-1_0.cube
3. Step
Each cube-file contains the electronic density of one MO mapped onto a regular 3D-grid. Not all MOs were written to a cube-file, this is controlled by the PRINT_MO section. Their filenames tell you to which MO a cube-file belongs. For example ethene-WFN_00005_1-1_0.cube contains the 5th orbital.
Use VMD to visualize the cube-files:
- To run:
$ vmd ethene-WFN_00008_1-1_0.cube - To visualize the molecule (sometimes it's not visible by default):
go to Graphics > Representations > Draw style and set Drawing Method to CPK - Add a second representation by clicking on Create Rep
- In this second representation set Drawing Method=Isosurfaces and Draw=Wireframe
- Finally set the Isovalue of to a reasonable value, eg. 0.1 .
- To visualize the positive and the negative part of an orbital simultaneously, you will have to add a third representation with a negative Isovalue, e.g. -0.1 .
- To give the two representations different colors, set their Coloring Method=ColorID and choose different ids.
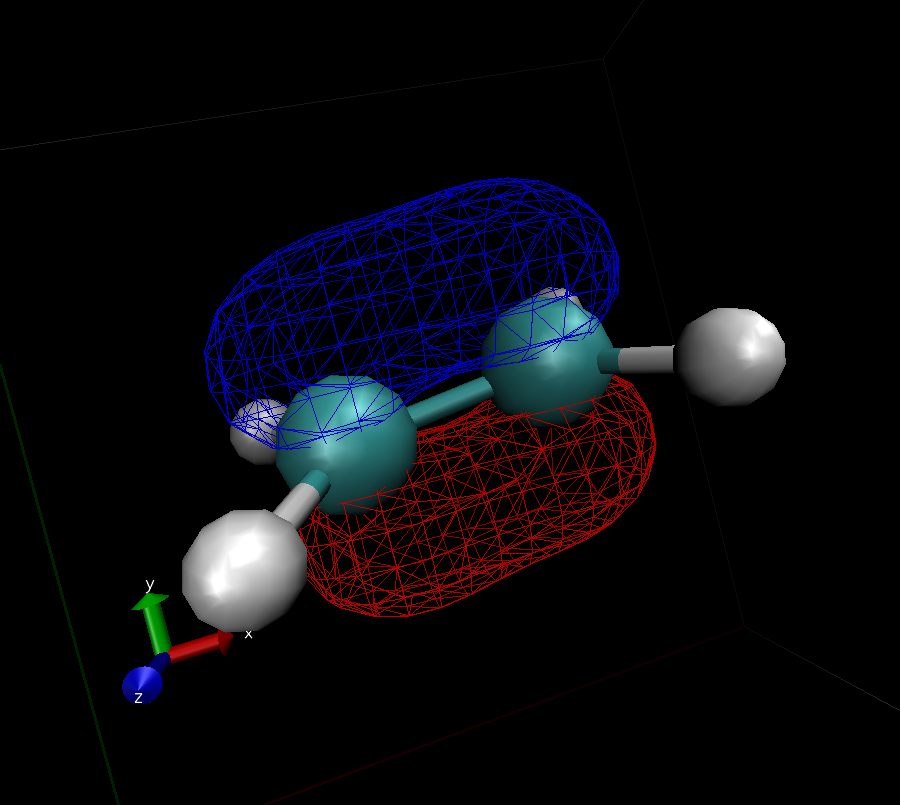
What you get should look similar to this:
Questions
- Compare the new input file with the one from the previous exercise: which keywords changed? which section is missing, respectively new? Lookup the description of the changed keywords and sections in the CP2K Manual
- From the output: What are the energies of the Highest Occupied MO (HOMO), Lowest Unoccupied MO (LUMO), and the band-gap (in electronvolt)?
- Use VMD to identify the shape of the $\pi$ and $\pi^*$ orbitals (submit images like the one from above)
- Repeat the calculation for Propene and find again the HOMO, LUMO and band-gap energies.
- The eigenvalues are given in Hartree (Eh) while the band-gap is stated directly in electronvolt
- Lookup the molecular orbital diagram of Ethen to identify which MOs and therefore which cube files you need to open.
- Use the Computational Chemistry Comparison and Benchmark DataBase to lookup the calculated geometry for Propene (CH2CHCH3), use the geometry made using a Hartree-Fock calculation and the
6-311+G(3df,2pd)basis set.