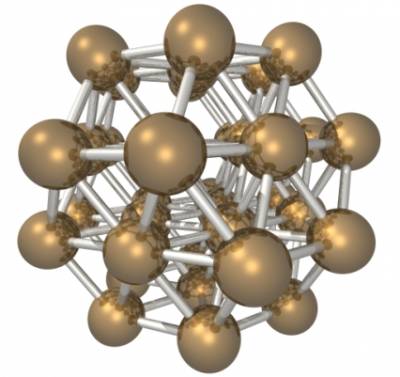
38 atom Lennard-Jones cluster
you@eulerX ~$ module load courses mmm vmd
you@eulerX ~$ mmm-init
you@eulerX ~$ module load cp2k
and to submit the job (note: since all the examples of this week are ultrafast, we will run them interactively, and NOT on a compute node. This is not the normal procedure for the next lectures).
you@eulerX ~$ cp2k.popt -i file.inp -o file.out
Download the 1.1 exercise into your $HOME folder and unzip it.
you@eulerX ~$ wget http://www.cp2k.org/_media/exercises:2017_ethz_mmm:exercise_1.1.zip you@eulerX ~$ unzip exercises:2017_ethz_mmm:exercise_1.1.zip
In this exercise you will test the Lennard-Jones potential. In particular, we will focus on the system described in the following paper about the energy landscape of the 38 atom Lennard-Jones cluster:
Login to euler using your nethz credentials. Then go to the directory “exercise_1.1”.
you@eulerX ~$ cd exercise_1.1
Geometry optimization
In this first part you will perform a simple energy optimization, to find the two lowest lying minima in the potential energy surface.
The input file structure of the template is the following:
- geo_opt.inp
&GLOBAL FLUSH_SHOULD_FLUSH PRINT_LEVEL low PROJECT geo_opt_bfgs RUN_TYPE geo_opt WALLTIME 600 &END GLOBAL &MOTION &GEO_OPT OPTIMIZER BFGS MAX_ITER 200 MAX_DR 0.001 RMS_DR 0.0003 MAX_FORCE 0.0001 RMS_FORCE 0.00003 &BFGS USE_MODEL_HESSIAN yes &END BFGS &END GEO_OPT &PRINT &TRAJECTORY on FORMAT xyz &EACH GEO_OPT 1 &END EACH &END TRAJECTORY &END PRINT &END MOTION &FORCE_EVAL METHOD Fist STRESS_TENSOR ANALYTICAL &MM &FORCEFIELD &CHARGE ATOM Ar CHARGE 0.0 &END &NONBONDED &LENNARD-JONES atoms Ar Ar EPSILON 119.8 SIGMA 3.405 RCUT 8.4 &END LENNARD-JONES &END NONBONDED &CHARGE ATOM Kr CHARGE 0.0 &END CHARGE &END FORCEFIELD &POISSON PERIODIC NONE &EWALD EWALD_TYPE none &END EWALD &END POISSON &PRINT &FF_INFO OFF SPLINE_DATA SPLINE_INFO &END FF_INFO &END PRINT &END MM &PRINT &FORCES off &END FORCES &GRID_INFORMATION &END GRID_INFORMATION &PROGRAM_RUN_INFO &EACH GEO_OPT 1 &END EACH &END PROGRAM_RUN_INFO &STRESS_TENSOR &EACH GEO_OPT 1 &END EACH &END STRESS_TENSOR &END PRINT &SUBSYS &CELL A 100 0 0 B 0 100 0 C 0 0 100 PERIODIC NONE &END CELL &TOPOLOGY COORD_FILE_NAME in.xyz COORDINATE xyz &END &PRINT &CELL &END CELL &KINDS &END KINDS &MOLECULES OFF &END MOLECULES &SYMMETRY &END SYMMETRY &END PRINT &END SUBSYS &END FORCE_EVAL
- load the module with the special m_* bash functions and initialize the module:
module load courses mmm ; mmm-init
- randomize the coordinate files fcc.xyz
m_xyzrand 1.0 < fcc.xyz > fcc_rand.xyz
. Do the same with ico.xyz
- extract the q4 order parameter from fcc.xyz and from fcc_rand.xyz and compare the values.
module load new gcc/4.8.2 python/2.7.12 python stein.py file.xyz
. You will be asked the cutoff radius for the neighbors, it is 1.391 in sigma units. You should input it in Angstrom.
- before running the simulation, copy the input coordinate file into in.xyz
cp fcc_rand.xyz in.xyz
- run cp2k
module load cp2k
(this has only to be done once)
cp2k.popt -i geo_opt.inp -o geo_opt.out
- in the output file, note the final energy, transform it in the unit of the paper (epsilon units)
- load vmd module and play with the optimization trajectory
vmd OPT-pos-1.xyz
(ask the teacher)
- apply the script myq4 to the optimization trajectory: this generates a list of q4 and energies for the whole trajectory.
./myq4 OPT-pos-1.xyz > fcc.ene.q4
- plot q4 and energies with gnuplot (ask the teacher)
- have a look at the myq4 script
nano myq4
- repeat for the ico.xyz starting point, don't forget to first copy/remove the files appropriately. For example:
mkdir FCC ; mv OPT* FCC ; mv geo_opt.out FCC
- finally, run the bash script
./curve
. Look inside, and try to understand what you get.
- Report the energy of the minima, compare it with the ones of the initial configurations.
- Plot q4 vs. energy and q4 vs. optimization steps, for the two cases. Discuss the results. Are the minima in two separate basins?
- Report the value of the order parameter of the minumum, and discuss what you see
- Use “gnuplot” to make the output of “./curve” understandable, discuss the results.