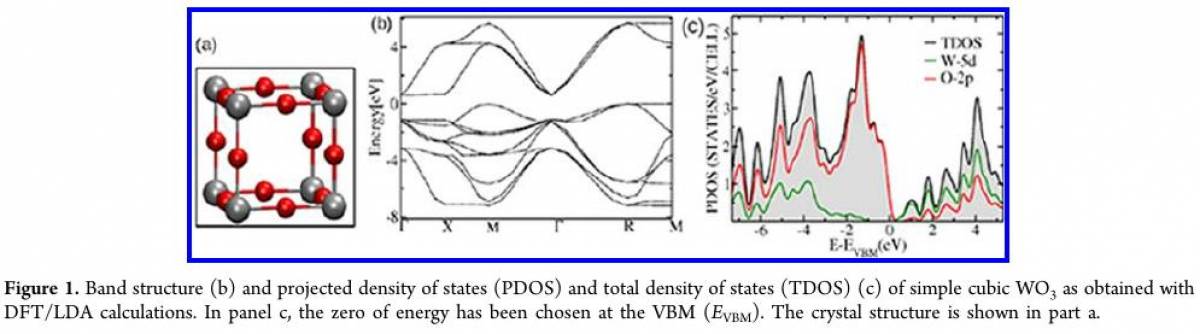
Projected density of states and Band structure for WO$_3$
In this exercise, you will carry out Density Of States(DOS) and band structure calculation using K-point sampling for Cubic lattice WO$_3$. The reference DOS and band structure you can find in this paper
Getting the PDOS
In the following exercise we are going to look at the density of states of WO3:
Similar to the previous exercise we write the coordinates in term of the unit cell:
- WO3-PDOS.inp
&GLOBAL PROJECT WO3-pdos RUN_TYPE ENERGY PRINT_LEVEL MEDIUM &END GLOBAL &FORCE_EVAL METHOD Quickstep &DFT BASIS_SET_FILE_NAME BASIS_MOLOPT POTENTIAL_FILE_NAME POTENTIAL &POISSON PERIODIC XYZ &END POISSON &SCF SCF_GUESS ATOMIC EPS_SCF 1.0E-6 MAX_SCF 300 ADDED_MOS 100 &DIAGONALIZATION ALGORITHM STANDARD EPS_ADAPT 0.01 &END DIAGONALIZATION &SMEAR ON METHOD FERMI_DIRAC ELECTRONIC_TEMPERATURE [K] 300 &END SMEAR &MIXING METHOD BROYDEN_MIXING ALPHA 0.2 BETA 1.5 NBROYDEN 8 &END MIXING &END SCF &XC &XC_FUNCTIONAL PBE &END XC_FUNCTIONAL &END XC &PRINT &PDOS # print all projected DOS available: NLUMO -1 # split the density by quantum number: COMPONENTS &END &END PRINT &END DFT &SUBSYS &CELL ABC [angstrom] 3.810000 3.810000 3.810000 PERIODIC XYZ MULTIPLE_UNIT_CELL 2 2 2 &END CELL &TOPOLOGY MULTIPLE_UNIT_CELL 2 2 2 &END TOPOLOGY &COORD SCALED W 0.0 0.0 0.0 O 0.5 0.0 0.0 O 0.0 0.5 0.0 O 0.0 0.0 0.5 &END &KIND W ELEMENT W BASIS_SET DZVP-MOLOPT-SR-GTH POTENTIAL GTH-PBE &END KIND &KIND O ELEMENT O BASIS_SET DZVP-MOLOPT-SR-GTH POTENTIAL GTH-PBE &END KIND &END SUBSYS &END FORCE_EVAL
The replication of the unit cell is necessary since the program samples only at the $\Gamma$ point unless instructed otherwise and we will otherwise do get a meaningful sampling of the density of states (e.g. the grid over the Brillouin Zone will be too coarse). Another option (which we will look into in the next exercise) is to sample over k-points instead.
What you will get in addition to the output file is a file named WO3_pdos-k1-1.pdos (to be precise, you will get one such file per atom kind but here we only have one, carbon) with a content similar to:
# Projected DOS for atomic kind W at iteration step i = 0, E(Fermi) = 0.144475 a.u.
# MO Eigenvalue [a.u.] Occupation s py pz px d-2 d-1 d0 d+1 d+2 f-3 f-2 f-1 f0 f+1 f+2 f+3
1 -2.621088 2.000000 0.87115225 0.00000000 0.00000000 0.00000000 0.00000000 0.00000000 0.00000256 0.00000000 0.00000256 0.00000000 0.00000000 0.00000000 0.00000000 0.00000000 0.00000000 0.00000000
2 -2.621080 2.000000 0.85006340 0.00131878 0.00166813 0.00134861 0.00000000 0.00000000 0.00515023 0.00000000 0.00441289 0.00006412 0.00000000 0.00003847 0.00012977 0.00003934 0.00000000 0.00006557
[...]
The columns correspond to the orbitals present in the basis set (hence projected DOS). You would now do a convolution plot using a gaussian to get a smooth DOS
To the convoluted DOS, you may want to check this website. Here, it is provided two Python script to do the convolution. Download two files pdos.py and get-smearing-pdos.py to your folder. And execute the Python script using
python get-smearing-pdos.py file.pdos
Alternatively, you could also use the Python script developed by Tiziano, the parser bug was fixed already.
Please also note the unit of the energy, it is in $E_h$. When looking at DOS plots you may want to convert it to Electronvolt instead. In the convolution program, this has been done in the code.
While some of the new options to help with convergence are of numerical nature, the smearing is not.
- Repeat the above calculation for the different multiple cells 3x3x3, 4x4x4
- Get the Total DOS and PDOS of O$_2p$ and W$_5d$ orbitals and compare to the literature value.
- Do you see why it is necessary to do the unit cell replication?
- What is the value of WO$_3$ band gap? Compare the plots for 3x3x3 and 4x4x4.
- .. which state ($s$, $p_x$, ..) is mainly responsible for that?
- Change the $\sigma$ value in convolution program, and determine a reasonable value for the PDOS plot
Getting the band structure of WO$_3$ Lattice
To get the band structure for WO3, only a few changes are required compared to the previous example for calculating the PDOS:
- WO3-bs.inp
&GLOBAL PROJECT WO3-kp-bs RUN_TYPE ENERGY PRINT_LEVEL MEDIUM &END GLOBAL &FORCE_EVAL METHOD Quickstep &DFT BASIS_SET_FILE_NAME BASIS_MOLOPT POTENTIAL_FILE_NAME POTENTIAL &POISSON PERIODIC XYZ &END POISSON &QS EXTRAPOLATION USE_GUESS ! required for K-Point sampling &END QS &SCF SCF_GUESS ATOMIC EPS_SCF 1.0E-6 MAX_SCF 300 ADDED_MOS 2 &DIAGONALIZATION ALGORITHM STANDARD EPS_ADAPT 0.01 &END DIAGONALIZATION &SMEAR ON METHOD FERMI_DIRAC ELECTRONIC_TEMPERATURE [K] 300 &END SMEAR &MIXING METHOD BROYDEN_MIXING ALPHA 0.2 BETA 1.5 NBROYDEN 8 &END MIXING &END SCF &XC &XC_FUNCTIONAL PBE &END XC_FUNCTIONAL &END XC &KPOINTS SCHEME MONKHORST-PACK 3 3 1 WAVEFUNCTIONS REAL SYMMETRY .FALSE. FULL_GRID .FALSE. PARALLEL_GROUP_SIZE -1 &END KPOINTS &PRINT &BAND_STRUCTURE ADDED_MOS 2 FILE_NAME WO3.bs &KPOINT_SET UNITS B_VECTOR SPECIAL_POINT ??? #GAMA SPECIAL_POINT ??? #X SPECIAL_POINT ??? #M SPECIAL_POINT ??? #GAMA SPECIAL_POINT ??? #R SPECIAL_POINT ??? #M NPOINTS ??? &END &END BAND_STRUCTURE &END PRINT &END DFT &SUBSYS &CELL ABC [angstrom] 3.810000 3.810000 3.810000 PERIODIC XYZ MULTIPLE_UNIT_CELL 1 1 1 &END CELL &TOPOLOGY MULTIPLE_UNIT_CELL 1 1 1 &END TOPOLOGY &COORD SCALED W 0.0 0.0 0.0 O 0.5 0.0 0.0 O 0.0 0.5 0.0 O 0.0 0.0 0.5 &END &KIND W ELEMENT W BASIS_SET DZVP-MOLOPT-SR-GTH POTENTIAL GTH-PBE &END KIND &KIND O ELEMENT O BASIS_SET DZVP-MOLOPT-SR-GTH POTENTIAL GTH-PBE &END KIND &END SUBSYS &END FORCE_EVAL
Some notes on the input file:
- By specifying the
KPOINTsection you are enabling the K-Point calculation. - While you could specify the K-Points directly, we are using the Monkhorst-Pack scheme 1) to generate them. The numbers following the keyword
MONKHORST-PACKspecify the tiling of the brillouin zone. - After the basic calculation, CP2K calculates the energies along certain lines, denoted as
KPOINT_SET(when you check the documentation you will note that this section can be repeated). - The keyword
NPOINTSspecifies how many points (in the addition to the starting point) should be sampled between two special points. - The
SPECIAL_POINTkeyword is used to specify the start-, mid- and endpoints of the line. Those points usually denote special points in the reciprocal lattice/unit cell, like the $\Gamma$, $M$ or $K$ point. You can find the definition for these in the appendix section of this paper. This keyword can also be specified multiple times, making it possible to directly get the band structure for a complete path.
- WO3-cubic.xyz
4 WO3; a=3.810000 W 0.000000 0.000000 0.000000 O 1.905000 0.000000 0.000000 O 0.000000 1.905000 0.000000 O 0.000000 0.000000 1.905000
Now, when you run this input file you will get in addition the the output file, a file named WO3.bs which will look similar to the following:
SET: 1 TOTAL POINTS: 26
POINT 1 ******** ******** ********
POINT 2 ******** ******** ********
POINT 3 ******** ******** ********
POINT 4 ******** ******** ********
POINT 5 ******** ******** ********
POINT 6 ******** ******** ********
Nr. 1 Spin 1 K-Point 0.00000000 0.00000000 0.00000000
20
-73.66652408 -38.53370023 -37.80464132 -37.79327769
-16.71308703 -16.11075946 -16.02553853 -1.43495530
-1.34739188 -1.33357408 0.37912017 0.38948689
0.39582882 0.40030859 0.46965212 0.47418816
2.60728842 2.62105342 3.16044140 6.99806305
Nr. 2 Spin 1 K-Point 0.00000000 0.10000000 0.00000000
20
-73.66647294 -38.53337818 -37.80859042 -37.79536623
-16.67479677 -16.09554462 -15.96731960 -1.68492873
-1.44087258 -1.34318045 0.09257368 0.13769271
0.21643888 0.38447849 0.44179002 0.45757924
2.61768501 3.02368022 3.51828287 7.06644645
[...]
For each set there is a block named SET with the special points listed as POINT, followed by sub-blocks for each K-Point containing the energies for each MO.
- Lookup the special points for the $\Gamma$, $X$,$M$,$R$ points in the mentioned paper (make sure you choose the right lattice). Calculate and plot the band structure for WO3 lattice along $\Gamma$, $X$,$M$,$\Gamma$,$R$,$M$ (you are free to decide whether to use multiple K-Point sets are multiple special points in a single set). Mark the special points. Choose an appropriate number of interpolation points to get a smooth plot.
- Compare your plot with plots from literature. What is different?
- How many orbital energies do you get and why? Try to change the input to get more unoccupied orbitals.
To convert the band structure file to a file which can be plotted directly, you can use the script cp2k_bs2csv.py from below, which when passed a band structure file WO3.bs as an argument will write files WO3.bs-set-1.csv for each set containing the K-Point coordinates and the energies in one line.
To plot the WO3.bs-set-1.csv file, you can either load it into $MATLAB$ or use $GNUPLOT$ command line.
gnuplot>set yrange [-8:14] gnuplot>plot for [i=4:23] "WO3.bs.set-1.csv" u 0:i w l t ""
- cp2k_bs2csv.py
#!/usr/bin/env python """ Convert the CP2K band structure output to CSV files """ import re import argparse SET_MATCH = re.compile(r''' [ ]* SET: [ ]* (?P<setnr>\d+) [ ]* TOTAL [ ] POINTS: [ ]* (?P<totalpoints>\d+) [ ]* \n (?P<content> [\s\S]*?(?=\n.*?[ ] SET|$) # match everything until next 'SET' or EOL ) ''', re.VERBOSE) SPOINTS_MATCH = re.compile(r''' [ ]* POINT [ ]+ (?P<pointnr>\d+) [ ]+ (?P<a>\S+) [ ]+ (?P<b>\S+) [ ]+ (?P<c>\S+) ''', re.VERBOSE) POINTS_MATCH = re.compile(r''' [ ]* Nr\. [ ]+ (?P<nr>\d+) [ ]+ Spin [ ]+ (?P<spin>\d+) [ ]+ K-Point [ ]+ (?P<a>\S+) [ ]+ (?P<b>\S+) [ ]+ (?P<c>\S+) [ ]* \n [ ]* (?P<npoints>\d+) [ ]* \n (?P<values> [\s\S]*?(?=\n.*?[ ] Nr|$) # match everything until next 'Nr.' or EOL ) ''', re.VERBOSE) if __name__ == '__main__': parser = argparse.ArgumentParser(description=__doc__) parser.add_argument('bsfilename', metavar='bandstructure-file', type=str, help="the band structure file generated by CP2K") args = parser.parse_args() with open(args.bsfilename, 'r') as fhandle: for kpoint_set in SET_MATCH.finditer(fhandle.read()): filename = "{}.set-{}.csv".format(args.bsfilename, kpoint_set.group('setnr')) set_content = kpoint_set.group('content') with open(filename, 'w') as csvout: print(("writing point set {}" " (total number of k-points: {totalpoints})" .format(filename, **kpoint_set.groupdict()))) print(" with the following special points:") for point in SPOINTS_MATCH.finditer(set_content): print(" {pointnr}: {a}/{b}/{c}".format( **point.groupdict())) for point in POINTS_MATCH.finditer(set_content): results = point.groupdict() results['values'] = " ".join(results['values'].split()) csvout.write("{a} {b} {c} {values}\n".format(**results))