Table of Contents
Basic electronic structure calculation
In this exercise, you will perform a first basic electronic structure calculation to obtain the molecular orbitals (MOs) of Ethene: Your calculations will produce a list of occupied and non occupied MOs and a series of *.cube files, that allow the visualization of the orbitals with VMD.
It is recommended to install and run VMD on your local machine. VMD can be downloaded free of charge after registering with your name and email address. To load and render the *.cube files on your local VMD you first have to transfer them from the server to your machine by using one of the transfer tools recommended on the First Login page.
1. Step: Run the calculation
Create a new directory for this exercise and run a CP2K calculation with the following (commented) input file (look at the first exercise again to figure out how to run a simulation):
- ethene.inp
&GLOBAL PROJECT ethene RUN_TYPE ENERGY PRINT_LEVEL MEDIUM &END GLOBAL &FORCE_EVAL METHOD Quickstep ! Electronic structure method (DFT,...) &DFT &PRINT &MO_CUBES ! Controls which MOs are written to cube-files. NHOMO 5 NLUMO 5 &END MO_CUBES &END PRINT &POISSON ! Solver requested for non periodic calculations PERIODIC NONE PSOLVER WAVELET ! Type of solver &END POISSON &QS ! Parameters needed to set up the Quickstep framework METHOD GAPW ! Method: gaussian and augmented plane waves &END QS &SCF ! Parameters controlling the convergence of the scf. This section should not be changed. MAX_ITER_LUMOS 10000 EPS_SCF 1.0E-6 SCF_GUESS ATOMIC MAX_SCF 60 EPS_LUMOS 0.000001 &OUTER_SCF EPS_SCF 1.0E-6 MAX_SCF 6 &END &END SCF &XC ! Parameters needed to compute the electronic exchange potential &XC_FUNCTIONAL NONE ! No xc functional &END XC_FUNCTIONAL &HF ! Hartree Fock exchange. In this case is 100% (no fraction specified). &SCREENING ! Screening of the electronic repulsion up to the given threshold. EPS_SCHWARZ 1.0E-10 ! Threshold specification &END SCREENING &END HF &END XC &END DFT &SUBSYS &CELL ABC 10 10 10 PERIODIC NONE ! Non-periodic calculations. That's why the POISSON section is needed &END CELL &TOPOLOGY ! Section used to center the atomic coordinates in the given box. Useful for big molecules &CENTER_COORDINATES &END &END &COORD C -2.15324 3.98235 0.00126 C -0.83403 4.16252 -0.00140 H -0.25355 3.95641 0.89185 H -0.33362 4.51626 -0.89682 H -2.65364 3.62861 0.89669 H -2.73371 4.18846 -0.89198 &END COORD &KIND H ! Basis set and potential for H &BASIS 2 1 0 0 3 1 18.73113700 0.03349460 2.82539370 0.23472695 0.64012170 0.81375733 1 0 0 1 1 0.16127780 1.00000000 &END POTENTIAL ALL &POTENTIAL 1 0 0 0.20000000 0 &END &END KIND &KIND C ! Basis set and potential for C &BASIS 4 1 0 0 6 1 3047.52490000 0.00183470 457.36951000 0.01403730 103.94869000 0.06884260 29.21015500 0.23218440 9.28666300 0.46794130 3.16392700 0.36231200 1 0 1 3 1 1 7.86827240 -0.11933240 0.06899910 1.88128850 -0.16085420 0.31642400 0.54424930 1.14345640 0.74430830 1 0 1 1 1 1 0.16871440 1.00000000 1.00000000 1 2 2 1 1 0.80000000 1.00000000 &END POTENTIAL ALL &POTENTIAL 4 2 0 0.34883045 0 &END &END KIND &END SUBSYS &END FORCE_EVAL
2. Step
If the calculation was performed correctly, a number of new files should have been written:
$ ls *.cube ethene-WFN_00004_1-1_0.cube ethene-WFN_00008_1-1_0.cube ethene-WFN_00012_1-1_0.cube ethene-WFN_00005_1-1_0.cube ethene-WFN_00009_1-1_0.cube ethene-WFN_00013_1-1_0.cube ethene-WFN_00006_1-1_0.cube ethene-WFN_00010_1-1_0.cube ethene-WFN_00007_1-1_0.cube ethene-WFN_00011_1-1_0.cube
3. Step
Each cube-file contains the electronic density of one MO mapped onto a regular 3D-grid. Not all MOs were written to a cube-file, this is controlled by the PRINT_MO section. Their filenames tell you to which MO a cube-file belongs. For example ethene-WFN_00005_1-1_0.cube contains the 5th orbital.
Use VMD to visualize the cube-files:
- Open one
.cubefile at a time in VMD - To visualize the molecule (sometimes it's not visible by default):
go to Graphics > Representations > Draw style and set Drawing Method to CPK - Add a second representation by clicking on Create Rep
- In this second representation set Drawing Method=Isosurfaces and Draw=Wireframe
- Finally set the Isovalue of to a reasonable value, eg. 0.1 .
- To visualize the positive and the negative part of an orbital simultaneously, you will have to add a third representation with a negative Isovalue, e.g. -0.1 .
- To give the two representations different colors, set their Coloring Method=ColorID and choose different ids.
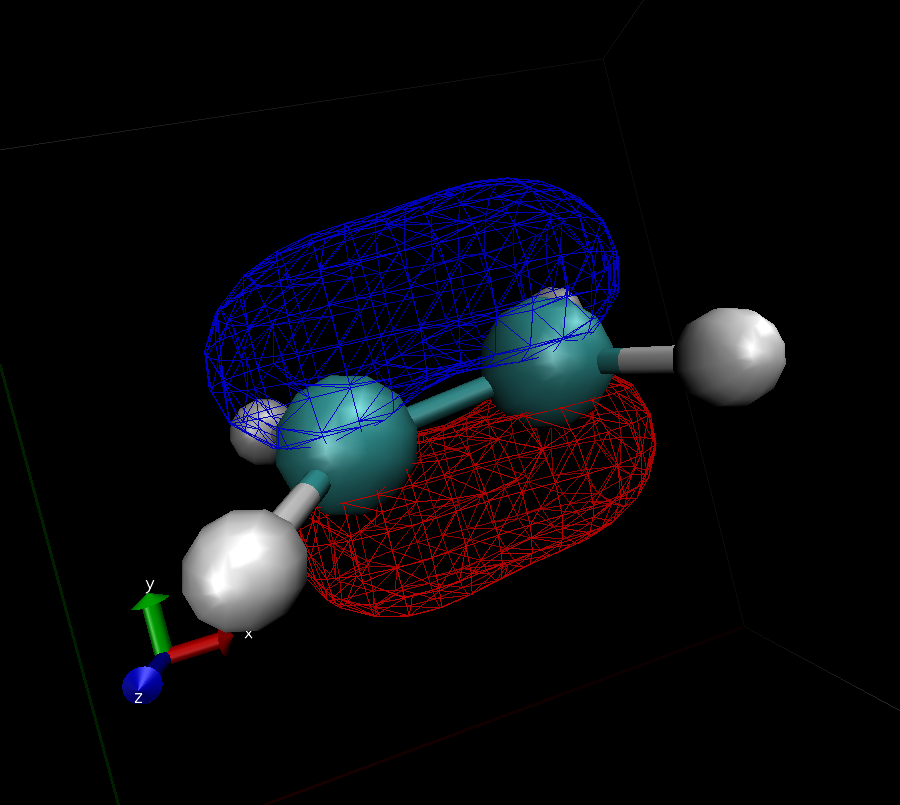
What you get should look similar to this:
Questions
- Compare the new input file with the one from the previous exercise: which keywords changed? which section is missing, respectively new? Lookup the description of the changed keywords and sections in the CP2K Manual
- From the output: What are the energies of the Highest Occupied MO (HOMO), Lowest Unoccupied MO (LUMO), and the band-gap (in electronvolt)?
- Use VMD to identify the shape of the $\pi$ and $\pi^*$ orbitals (submit images like the one from above)
- Repeat the calculation for Propene and find again the HOMO, LUMO and band-gap energies.
- The eigenvalues are given in Hartree (Eh) while the band-gap is stated directly in electronvolt
- Lookup the molecular orbital diagram of Ethen to identify which MOs and therefore which cube files you need to open.
- Use the Computational Chemistry Comparison and Benchmark DataBase to lookup the calculated geometry for Propene (CH2CHCH3), use the geometry optimized using a Hartree-Fock calculation and the
6-311+G(3df,2pd)basis set.