Table of Contents
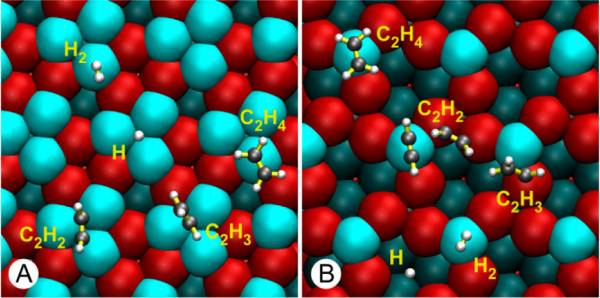
Adsorption of C2H2 on PdGa(111)
In this exercise you will compute the adsorption energy of acetylene on a intermetallic catalyst. This process is important during the production of polyethylene, and the system is described in this paper: 10.1021%2Fja505936b.
- In the first part of the exercise you will consider the optimized configuration (already in the directory) and study the pure electronic adsorption energy, namely the difference between the total energy of the surface-molecule system and the energy of the molecule alone and surface alone in the same geometry as the surface-molecule system minimum structure. This will allow to show the binding pattern of the electronic density.
- In the second part, you will optimize the surface and the molecule separately; this will allow to compute the total adsorption energy.
1. Task: Familiarize yourself
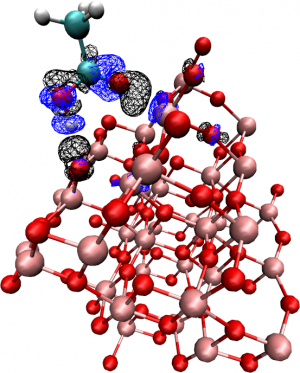
The coordinates of the optimized configuration are provided to you as S_M.opt.xyz (S stands for “Substrate”, M for “Molecule”, opt for “optimized”). Visualize the geometry with VMD and familiarize yourself with the system.
2. Task: Bond induced density differences
Compute the density difference induced by the adsorption bonding. For this you will have to run three separate energy calculations, using the *.ene.inp files.
- combined system (file
S_M.opt.xyz) - lone acetylene (file
M.S_M.xyz) - lone slab (file
S.S_M.xyz)
In order to output the electronic densities as cube files, your input file has to contain the following snipped:
&DFT
&PRINT
&E_DENSITY_CUBE
&END E_DENSITY_CUBE
&END
&END DFT
qsub run -v INP=prefix. Check the run file for the number of nodes.
To process the cube files we are going to use the cubecruncher tool. It is part of CP2K and is in your exercise directory.
you@eulerX ~$ ./cubecruncher.x -i S_M-ELECTRON_DENSITY-1_0.cube -subtract S-ELECTRON_DENSITY-1_0.cube -o tmp.cube you@eulerX ~$ ./cubecruncher.x -i tmp.cube -subtract M-ELECTRON_DENSITY-1_0.cube -o Delta_ads.cube
The generated cube file is not aligned with the simulation cell. Center the cube file with the cubecruncher.x tool:
you@eulerX ~$ ./cubecruncher.x -center geo -i Delta_ads.cube -o Delta_ads-centered.cube
You can visualize the resulting file delta_ads-centered.cube with VMD. This has been covered in a previous exercise.
3. Task: Bonding energies
Compute the binding energy:
\[ E_\text{binding}=\sum E_\text{products} - \sum E_\text{reactants} \]
For this you will need the energy values of three systems:
- lone acetylene molecule (run geometry optimization, use energy of last step)
- lone slab (you can use the already geometry optimized coordinates from
S.opt.xyzat the end of the exercise) - combined system adsorbed (can be reused from previous task)
Questions
- Sketch briefly the geometry of the molecule when adsorbed and in the gas phase.
- Report the system energy for the bonded system, lone slab, and lone molecule.
- Can you estimate the contribution due to the geometry relaxation?
- Briefly report the bond induced density difference on the system.
Required Files
/home/psd/Exercise_9. Change the name of the xyz file accordingly in the input files.
- S_M.inp
&FORCE_EVAL METHOD Quickstep &DFT &PRINT &E_DENSITY_CUBE &END E_DENSITY_CUBE &END BASIS_SET_FILE_NAME ./BR POTENTIAL_FILE_NAME ./GR &QS EPS_DEFAULT 1.0E-10 METHOD GPW EXTRAPOLATION ASPC EXTRAPOLATION_ORDER 3 &END QS &MGRID CUTOFF 400 NGRIDS 5 &END &SCF MAX_SCF 20 SCF_GUESS RESTART EPS_SCF 1.0E-5 &OT PRECONDITIONER FULL_SINGLE_INVERSE MINIMIZER CG &END &OUTER_SCF MAX_SCF 50 EPS_SCF 1.0E-5 &END &PRINT &RESTART &EACH GEO_OPT 2 &END ADD_LAST NUMERIC FILENAME RESTART &END &RESTART_HISTORY OFF &END &END &END SCF &XC &XC_FUNCTIONAL PBE &END XC_FUNCTIONAL &END XC &END DFT &SUBSYS &CELL A [angstrom] 14.08557 0 0 B [angstrom] 0 12.1985 0 C [angstrom] 0.000000 0.000000 15.0 &END CELL &TOPOLOGY COORD_FILE_NAME ./S_M.opt.xyz COORDINATE xyz &END &KIND Pd BASIS_SET DZVP-MOLOPT-SR-GTH-q18 POTENTIAL GTH-PBE-q18 &END KIND &KIND Ga BASIS_SET DZVP-MOLOPT-SR-GTH-q13 POTENTIAL GTH-PBE-q13 &END KIND &KIND C BASIS_SET TZV2P-MOLOPT-GTH POTENTIAL GTH-PBE-q4 &END KIND &KIND H BASIS_SET TZV2P-MOLOPT-GTH POTENTIAL GTH-PBE-q1 &END KIND &END SUBSYS &END FORCE_EVAL &GLOBAL PRINT_LEVEL LOW PROJECT S_M RUN_TYPE ENERGY &END GLOBAL