Table of Contents
Protein Folding in Solution
In this exercise, you will calculate the protein folding free energy using thermodynamic integration, a method based on molecular dynamics (MD). The protein will be described by the empirical force field, CHARMM22, http://mackerell.umaryland.edu/charmm_ff.shtml
Background
A model protein you will have to deal with is the alanine decapeptide. The folding/unfolding will be achieved by stretching/compressing the chain. This in practice can be achieved by constraining the distance between the end carbon atoms in the protein and performing different simulations for each value of the distance. The atoms are the number 11 and 91, which are the carbon atoms of the carbonyl groups at the edges of the protein. The distance between these atoms is the collective variable chosen for the system. At each distance one runs a constrained MD to extract the time-averaged forces acting on the collective variable, $F(x)$. Then, a free energy difference can be calculated via thermodynamic integration (TI):
\begin{equation} \Delta A = -\int_a^b F(x)\, dx \end{equation}
Here $a$ and $b$ are the initial and the final values of the collective variable. TI is a general method, which can be applied to a variety of processes, e.g. phase transitions, electron transfer etc.
Task 1: Familiarize yourself
Download the files: deca_ala_ex3.tar.gz
in the directory deca_ala you will find
deca_ala.pdb (protein data base) file contains the coordinates
deca_ala.psf (protein structure file) file contains connectivity data
par_all27_prot_lipid.inp contains the force field parameters. You will be using the CHARMM v.22, a popular force field for biologically relevant systems.
md_std.inp is the CP2K input file
Open the deca_ala.pdb protein data bank format file with vmd. Create a new representation for the protein, e.g. of type Ribbon to observe the alpha-helix.
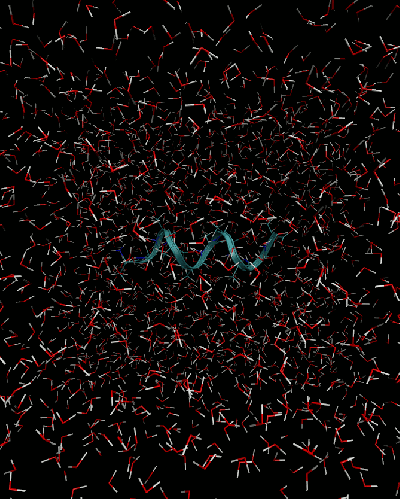
Although the image below shows the deca-alanine in water, it is expensive to run thermodynamic integration for a solvated protein with many values of the constraints on small laptops. So we will run TI for the protein in the gas-phase.
Task 2: Perform constrained MD simulations
Here you are asked to run several MD simulations for different values of the distance between atoms 11 and 91, in each run it will be constrained. In the original file md_std.inp the distance is set to $14.37$ Å as is in the deca_ala.pdb file. This is the first step to carry out the termodynamic integration, as described in the equation above.
We have made the script run_ti_jobs.sh to run these simulations, which you can find inside the compressed file deca_ala.tar.gz. Take a look at the script and familiarize yourself with it. At which values are we constraining the distances between the carbon atoms? In this case we are performing 5 different simulations, each with a different value of the constraint. You can edit this script to use a larger or smaller number of constraints and to increase or reduce the upper and/or lower bound of integration. Can you guess where in the script we are specifying the values of the constraints?
- run_ti_jobs.sh
#!/bin/bash -l for d in $(seq 16 1 20); do mkdir $d cp deca_ala/* $d/. cd $d sed -e "s|14.37|${d}|" md_std.inp > d_${d}.inp cp2k.sopt -i d_${d}.inp -o d_${d}.out cd .. done
- Be careful with the values chosen for the upper and lower bound of the constraints as the simulations might crash or the SHAKE algorithm for the computation of the constraints might not converge if the values of the constrained distances are unphysical.
- We have set the number of steps of each constrained MD to 5000. Try to increase this number if you want to achieve better statistics or to decrease it to get the results faster, at the expense of a more converged free energy.
Relevant constraint sections for constrained MD
Look into the main input file of cp2k md_std.inp, and try to understand the keywords used as much as possible, by now you should be able to understand most of it, and you can experiment changing some of the keywords to see what happens. Look in particular at the definition of the section CONSTRAINT where the target value of the distance between the two carbon atoms at the edges of the proteins are constrained for instance to 14.37, and at the COLVAR section where the the collective variable for the distance between the two C atoms is defined.
- constraint section
&CONSTRAINT &COLLECTIVE COLVAR 1 INTERMOLECULAR TARGET [angstrom] 14.37 &END COLLECTIVE &LAGRANGE_MULTIPLIERS COMMON_ITERATION_LEVELS 1 &END &END CONSTRAINT
- colvar section
&COLVAR &DISTANCE ATOMS 11 91 &END DISTANCE &END
Task 3: Evaluate the free energy difference
⇒ Each constrained MD will produce a .LagrangeMultLog-files, which look like this:
Shake Lagrangian Multipliers: -63.547262596 Rattle Lagrangian Multipliers: 63.240598387 Shake Lagrangian Multipliers: -0.326901815 Rattle Lagrangian Multipliers: -0.318145579
- From these files you can calculate the average Lagrange multiplier of the Shake-algorithm and the standard error like this:
grep Shake yourprojectname.LagrangeMultLog | awk '{c++ ; s=s+$4; sq=sq+$4*$4 }END{print s/c, sqrt(sq/c - s*s/(c*c))/(sqrt(c)) }'
- The average Lagrange multiplier is the average force $F(x)$ required to constrain the atoms at the distance $x$. First of all, plot the force $F(x)$ with its standard error as a function of the collective variable to see if the simulation carried out so far is statistically relevant or the relative error is too large.
- From the forces, the free energy difference can be obtained via thermodynamic integration between the two states. Given that state $a$ and $b$ are the initial and the final values of the collective variable, extract the free energy difference from
\begin{equation} \Delta A = -\int_a^b F(x)\, dx \end{equation}
- Also, extract the free energy profile as a function of the collective variable $x^\prime$. To do this, one can choose the minimum value of the collective variable, as the lower bound of the integral and plot the free energy as a function of $x^\prime$, the upper bound of the integral, according to the same relation
\begin{equation} \Delta A(x^\prime) = -\int_a^{x^\prime} F(x)\, dx \end{equation}
- Discuss the form of the free energy profile and comment on what is the most stable state of the protein. Is it more stable when it is stretched or when it is in the $\alpha$-helix conformation? Is this result physical? Explain why or why not. How can the presence of water affect the conformation of the protein?
- Tip 1: the most stable state will be that where the free energy is at the global minimum.
- Tip 2: In order to understand whether the result obtained from thermodynamic integration is physical or not, have a look at the
.xyzfiles for some of the constrained MD trajectories and think about what are the fundamental interactions between the constituents of the protein that we are taking into account with the CHARMM force field (e.g. electrostatic, van-der Waals, covalent bonds) and how these may contribute to the stabilization of the protein in a given state. - The two articles at the links below show how the free energy profile should look like, using thermodynamic integration or a different enhanced sampling method. Compare the free energy profile obtained from your simulations to either of those papers. Most likely, the free energy profile you obtained will not be as converged as theirs. What are some possible reasons for this, and how can one obtain better converged free energy profiles?
- Paper 1: https://arxiv.org/pdf/0711.2726.pdf see figure 2, solid line obtained with thermodynamic integration, using the same force field (CHARMM v.22) used here. This paper however, uses a different collective variable, i.e. the distance between the N-atoms at the opposite edges.
- Paper 2: https://pubs.acs.org/doi/pdf/10.1021/ct5002076 see figure 1, obtained with umbrella sampling and adaptive bias force sampling, for two versions of the CHARMM force field, v.22 and v.36. The collective variable in this case is the same as the one specified in our input.
- Finally, in principle we could have performed a direct MD simulation (as we did in the past exercises) to compute the free energy profile as a function of the distance between two of the atoms at the opposite edges of the protein (the collective variable we chose for this particular problem). Instead, we chose to perform an enhanced simulation technique. Can you think of a problem we would face if we had decided to perform a direct MD simulation? What could be a possible way to overcome this problem?
- We have provided you with a useful script called
generate_plots.shthat extracts the average force and the standard error for each constrained MD simulation (see thegrepcommand line above), and it prints out the fileav_force_vs_x.datcontaining the force as a function of the collective variable, and the error on the force (third column). Take a look at the script and modify it if necessary, e.g. if you have changed the lower and upper bound for the constraint or if you have changed the number of constraints. - In order to check the convergence of the free energy profile one should look at the error on the average force for each constrained MD simulation. The error on the free energy profile can be obtained by propagating the error on the average force upon integration.
- From the file containing the average force as a function of collective variable you need to integrate $F(x) dx$ numerically to obtain $\Delta A$. You may use the trapezoidal rule (or equivalent) with EXCEL, ORIGIN or any scripting language.